Master Scikit-learn Pipeline #
“You clean data. Then you scale it. Then you encode categories. Then you select features. Then you train a model. Then you realize you forgot to apply the same steps to your test data. You cry. Pipeline fixes this. It chains all your steps into one clean workflow.”
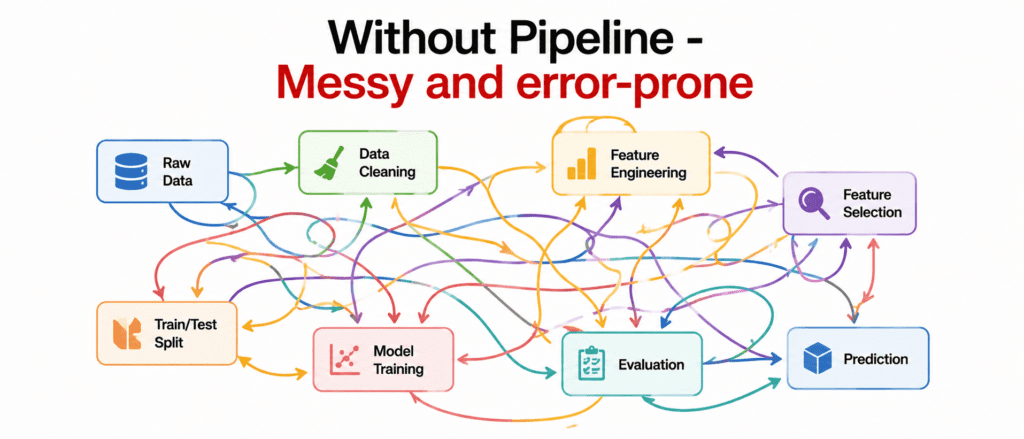
The Problem Without Pipeline #
You write code like this:
# Step 1: Scale scaler.fit(X_train) X_train_scaled = scaler.transform(X_train) X_test_scaled = scaler.transform(X_test) # Step 2: Encode encoder.fit(X_train_scaled) X_train_encoded = encoder.transform(X_train_scaled) X_test_encoded = encoder.transform(X_test_scaled) # Step 3: Select features selector.fit(X_train_encoded, y_train) X_train_selected = selector.transform(X_train_encoded) X_test_selected = selector.transform(X_test_encoded) # Step 4: Train model model.fit(X_train_selected, y_train)
Problems:
- You forget a step
- You transform in wrong order
- You leak data from test to train
- Code is messy and hard to share
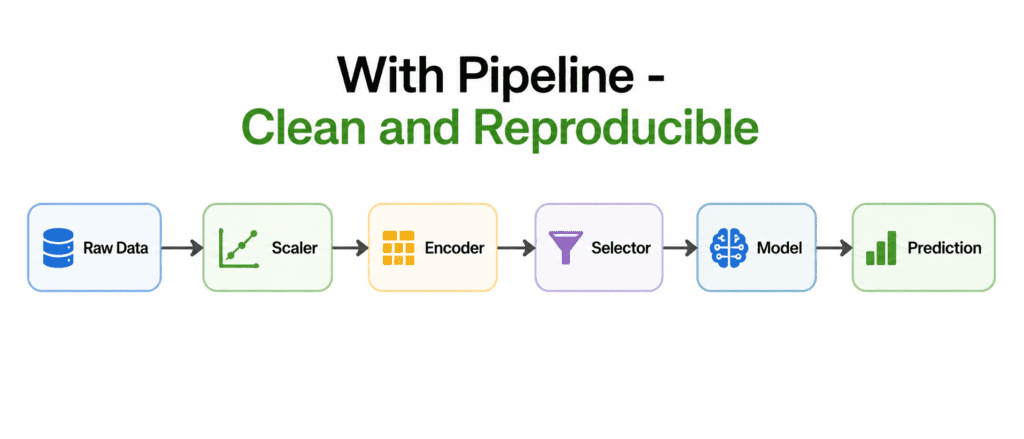
The Solution: Pipeline #
What is a Pipeline? A chain that connects preprocessing steps and final model. Data flows through each step in order.
How it looks:
from sklearn.pipeline import Pipeline
pipeline = Pipeline([
('scaler', StandardScaler()),
('encoder', OneHotEncoder()),
('selector', SelectKBest(k=10)),
('model', RandomForestClassifier())
])
# One line fits everything
pipeline.fit(X_train, y_train)
# One line predicts
predictions = pipeline.predict(X_test)Benefits:
- All steps in one object
- Same transformations apply to train and test automatically
- No data leakage
- Easy to share and deploy
- Easy to try different steps
How Pipeline Works #
When you call fit():
- Pipeline calls fit() on first step (scaler)
- Then transform() on first step
- Passes transformed data to second step
- Repeats through all steps
- Finally calls fit() on last step (model)
When you call predict():
- Pipeline calls transform() on all preprocessing steps
- Passes transformed data to final model
- Model predicts
Simple visual:
fit() flow: X → Step1.fit() → Step1.transform() → Step2.fit() → ... → Model.fit() predict() flow: X → Step1.transform() → Step2.transform() → ... → Model.predict()
Building Your First Pipeline #
Step 1: Import required pieces
from sklearn.pipeline import Pipeline from sklearn.preprocessing import StandardScaler from sklearn.decomposition import PCA from sklearn.ensemble import RandomForestClassifier
Step 2: Define the pipeline
pipeline = Pipeline([
('scaler', StandardScaler()), # Step 1: Scale features
('dim_reduction', PCA(n_components=50)), # Step 2: Reduce dimensions
('classifier', RandomForestClassifier()) # Step 3: Train model
])Step 3: Use it like any model
pipeline.fit(X_train, y_train) accuracy = pipeline.score(X_test, y_test) predictions = pipeline.predict(X_new)
The names (‘scaler’, ‘dim_reduction’, ‘classifier’) are for you. You can name them anything.
ColumnTransformer (Handling Mixed Data Types) #
Problem: Different columns need different preprocessing. Numeric columns need scaling. Categorical columns need encoding.
Solution: ColumnTransformer applies different pipelines to different columns.
Example:
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
# Define column groups
numeric_features = ['age', 'income', 'years_experience']
categorical_features = ['city', 'education', 'department']
# Create preprocessor
preprocessor = ColumnTransformer([
('numeric', StandardScaler(), numeric_features),
('categorical', OneHotEncoder(), categorical_features)
])
# Create full pipeline
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', RandomForestClassifier())
])
# Use it
pipeline.fit(X_train, y_train)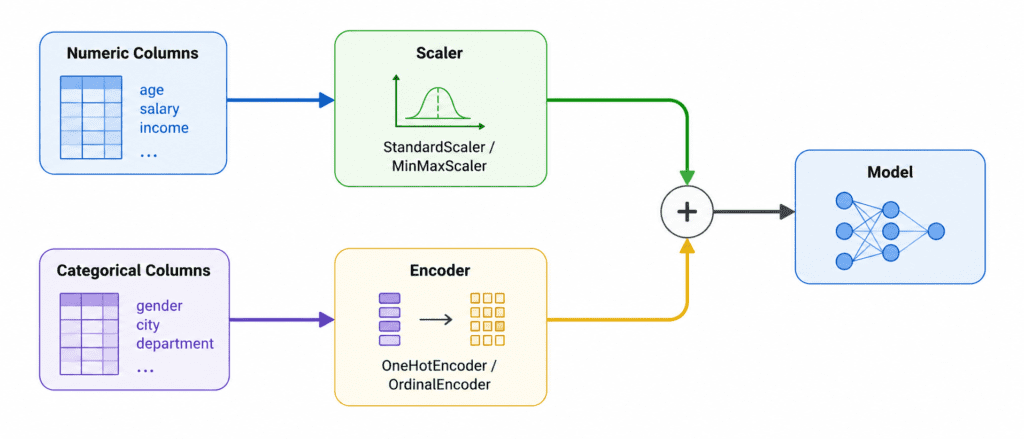
Real Example: California Housing Pipeline #
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestRegressor
# Define columns
numeric_cols = ['longitude', 'latitude', 'housing_median_age', 'total_rooms',
'total_bedrooms', 'population', 'households', 'median_income']
categorical_cols = ['ocean_proximity']
# Numeric pipeline
numeric_pipeline = Pipeline([
('imputer', SimpleImputer(strategy='median')),
('scaler', StandardScaler())
])
# Categorical pipeline
categorical_pipeline = Pipeline([
('imputer', SimpleImputer(strategy='most_frequent')),
('onehot', OneHotEncoder())
])
# Combine
preprocessor = ColumnTransformer([
('numeric', numeric_pipeline, numeric_cols),
('categorical', categorical_pipeline, categorical_cols)
])
# Full pipeline
full_pipeline = Pipeline([
('preprocessor', preprocessor),
('regressor', RandomForestRegressor(n_estimators=100))
])
# Train
full_pipeline.fit(X_train, y_train)
# Evaluate
score = full_pipeline.score(X_test, y_test)Pipeline with Grid Search (Hyperparameter Tuning) #
Pipeline works perfectly with GridSearchCV. You just add step names with double underscore.
Syntax: step_name__parameter_name
from sklearn.model_selection import GridSearchCV
pipeline = Pipeline([
('scaler', StandardScaler()),
('model', RandomForestClassifier(random_state=42))
])
# Define grid
param_grid = {
'model__n_estimators': [50, 100, 200],
'model__max_depth': [None, 10, 20],
'scaler__with_mean': [True, False] # Scaler also has parameters!
}
# Search
grid_search = GridSearchCV(pipeline, param_grid, cv=5)
grid_search.fit(X_train, y_train)
print(grid_search.best_params_)Notice: model__n_estimators not just n_estimators. The step name is required.
Pipeline with Different Models (Easy Comparison) #
Swap models without changing the rest:
# Same preprocessing, different models
pipelines = {
'rf': Pipeline([('preprocessor', preprocessor), ('model', RandomForestClassifier())]),
'svm': Pipeline([('preprocessor', preprocessor), ('model', SVC())]),
'lr': Pipeline([('preprocessor', preprocessor), ('model', LogisticRegression())])
}
# Compare
for name, pipeline in pipelines.items():
pipeline.fit(X_train, y_train)
score = pipeline.score(X_test, y_test)
print(f"{name}: {score:.4f}")Important Rules #
| # | Rule |
|---|---|
| 1 | Never fit pipeline on test data. Call fit() on train only. |
| 2 | Last step can be anything (classifier, regressor, or even transformer). |
| 3 | Step names must be unique. Cannot have two steps with same name. |
| 4 | Use ColumnTransformer when different columns need different handling. |
| 5 | Save the pipeline after training. It contains all preprocessing steps. |
Saving and Loading Pipeline #
import joblib
# Save
joblib.dump(pipeline, 'my_pipeline.pkl')
# Load
loaded_pipeline = joblib.load('my_pipeline.pkl')
# Use loaded pipeline directly
predictions = loaded_pipeline.predict(X_new)Magic: The loaded pipeline remembers everything. No need to reapply preprocessing manually.

Quick Quiz #
Q1: You have a pipeline with scaler then SVM. You call pipeline.fit(X_train, y_train). What happens to the scaler?
A1: Scaler fits on X_train (calculates mean and std), then transforms X_train. Transformed data goes to SVM.
Q2: You call pipeline.predict(X_test). Does scaler fit again?
A2: No. Scaler uses the mean and std from training data to transform X_test. No refitting.
Q3: You want to try different values for n_estimators in RandomForest. How do you write this in GridSearchCV?
A3: 'model__n_estimators': [50, 100, 200] where ‘model’ is the step name.
Q4: You have numeric and categorical columns. What should you use?
A4: ColumnTransformer with separate pipelines for each column type.
Q5: You saved your pipeline after training. You load it on another computer. Do you need the preprocessing code again?
A5: No. The loaded pipeline contains everything. Just call predict().
